Human Pangenome Reference Consortium
@HumanPangenome
Diverse Human References Drive Genomic Discoveries for Everyone
ID:1177246141144096769
26-09-2019 15:39:00
375 Tweets
3,0K Followers
2,2K Following
Follow People

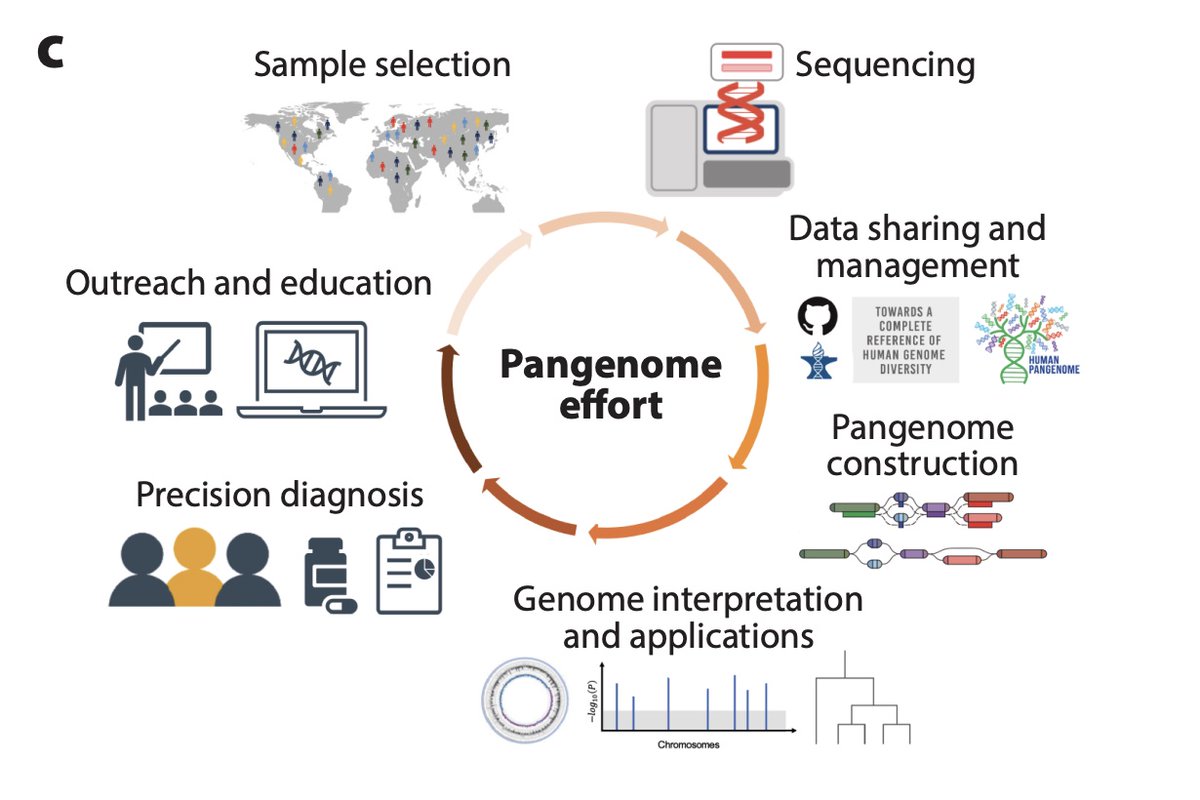
Excited to announce our new review: Beyond the Human Genome Project: The Age of Complete Human Genome Sequences and Pangenome References. w/ Benedict Paten Rajiv McCoy Karen Miga Eimear Kenny et al. annualreviews.org/content/journa…

Our review paper on 'Genome assembly in the telomere-to-telomere era' published online in Nature Reviews Genetics. It amazes me how much assembly has progressed in the past four years – often ~100X improvement in contiguity and base accuracy and with haplotypes and hard regions resolved!







Delighted to announce my friends and I are organising a 'Human Pangenome Variability' symposium at the #SMBE2024 in Puerto Vallarta 🇲🇽. We have two excellent invited speakers Arang Rhie and Erik Garrison. Submit your abstract by March 15th! All info at smbe2024.org



Memphis Pangenomics days, 2024 edition! Pangenome Workshop (May 18-19), Conference (May 20), & Biohackathon (May 21-22). pangenome.github.io/MemPanG24/ #Pangenomics #Bioinformatics #MemPanG24



Preprint on Exploring gene content with pangenome gene graphs: arxiv.org/abs/2402.16185. It describes pangene for building gene graphs and for calling gene-level variations which can be found at pangene.bioinweb.org.
Pleasant collaboration with Maximillian Marin and Maha Farhat.

#Pangenome graphs improve the analysis of SVs for rare genetic diseases. Our new study is out!
Great collaboration with the lab of Tomi Pastinen and congrats to Cristian Groza @[email protected] for the amazing work!
nature.com/articles/s4146…

We are proud to announce a new hg38 track group consisting of Human Pangenome Reference Consortium data. In this first release, there are tracks displaying features and variations of HPRC results compared to hg38, as well as alignment tracks.
See the full announcement: bit.ly/HPRCdataInUCSC…
















