Heng Li
@lh3lh3
Associate professor of biomedical informatics at @HarvardDBMI and @dfcidatascience
ID:2165843324
http://liheng.org 31-10-2013 02:21:20
3,7K Tweets
10,0K Followers
208 Following

Our new pre-print is live! 🧬🦠🎉
We performed a deep-sequencing experiment using PacBio HiFi reads (Revio & Sequel IIe) on a pooled human gut reference from Zymo Research, with a primary focus on #metagenome assembly.
1/6
doi.org/10.1101/2024.0…




We have released some raw Oxford Nanopore data for benchmarking purposes, along with some basecalled outputs.
We use the subset of 20k or 500k for benchmarking tools and methods a LOT!
We thought they would be helpful to others too.
4/5kHz R10.4.1 & RNA004
gentechgp.github.io/gtgseq/







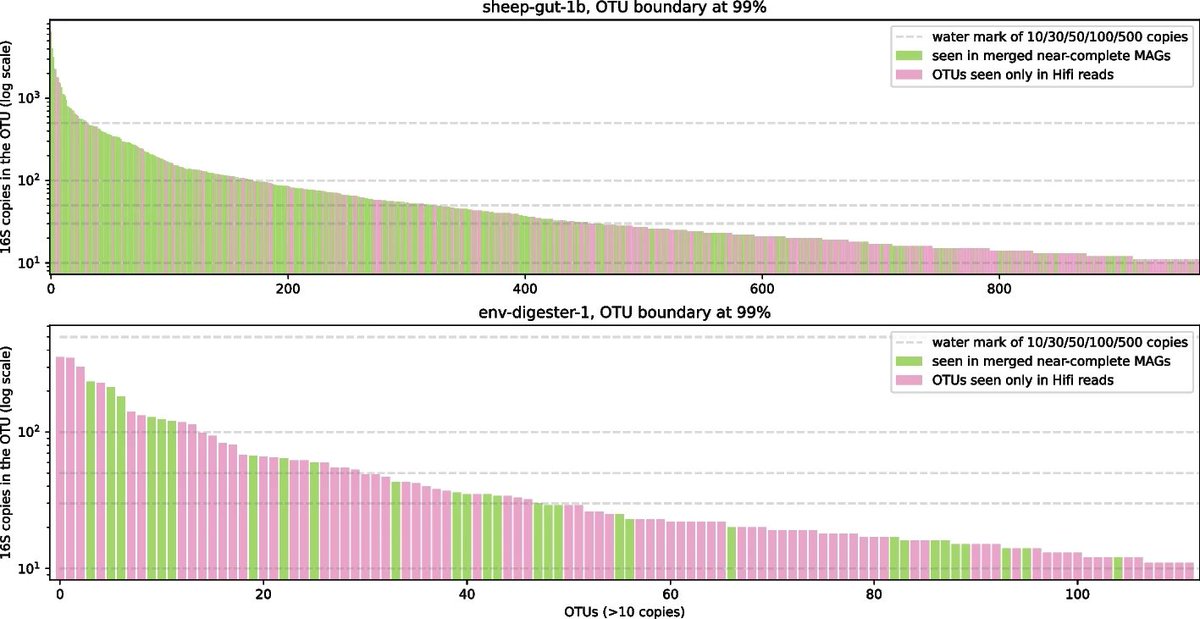
Xiaowen's (Xiaowen Feng) new paper uses 16S in reads and k-mers to answer how much a metagenome assembly captures the species and sequence content in a sample. We found abundant species are not always assembled and proposed a heuristic to rescue some of them. genomebiology.biomedcentral.com/articles/10.11…


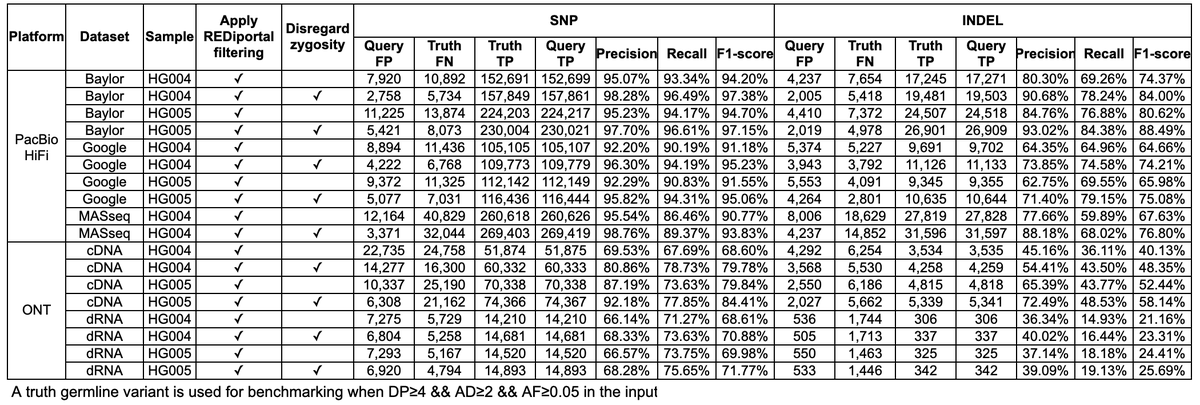
Making Clair3-RNA available, which is a side project of my lab inspired by Fritz Sedlazeck and Miten Jain last December. It's for RNA long-read germline variant calling. It supports ONT r9 cDNA, ONT rna002 dRNA, PB Sequel2 Iso-Seq, PB Revio MAS-Seq. 🔗github.com/HKU-BAL/Clair3… 1/n




Analysis of the limited M. tuberculosis accessory genome reveals potential pitfalls of pan-genome analysis approaches biorxiv.org/cgi/content/sh… #biorxiv_bioinfo